Gene phyloprofile
This interface allows the user to search for common OR specific genes/regions between a query genome and other genomes or replicons chosen from the ones available in our PkGDB database (i.e, (re)annotation of bacterial genomes) or complete proteome downloaded from the RefSeq/WGS sections.
How to read the interface?
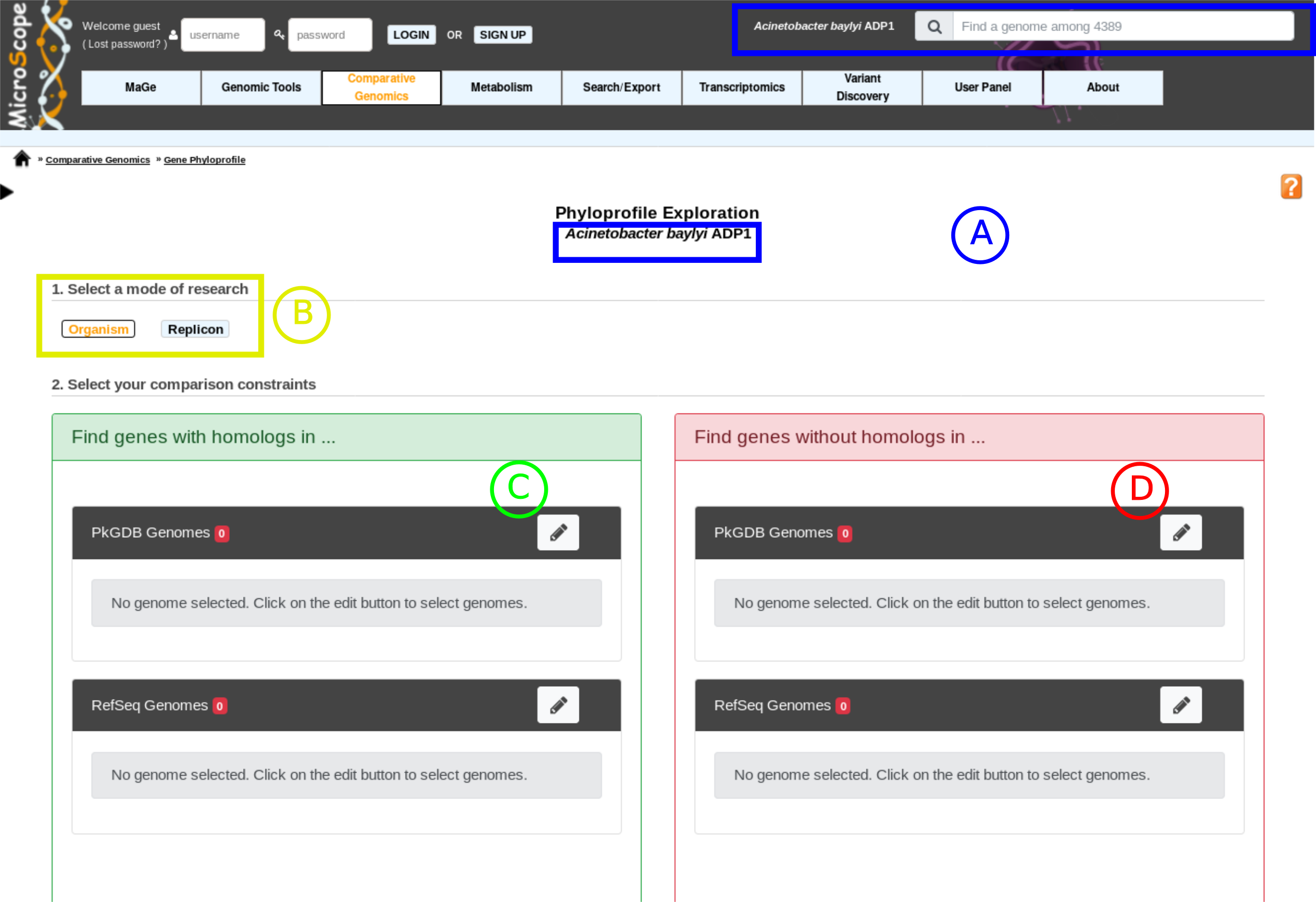
item A: Use the «Change» button to set the reference genome that will be used for the comparison. The current reference genome is displayed as a subtitle at the top of the window.
item B: Use this box to select the mode of comparison
in Organism mode, search is performed within all replicons of the selected organisms
in Replicon mode, search is performed within a specific replicon (chromosome/plasmid)
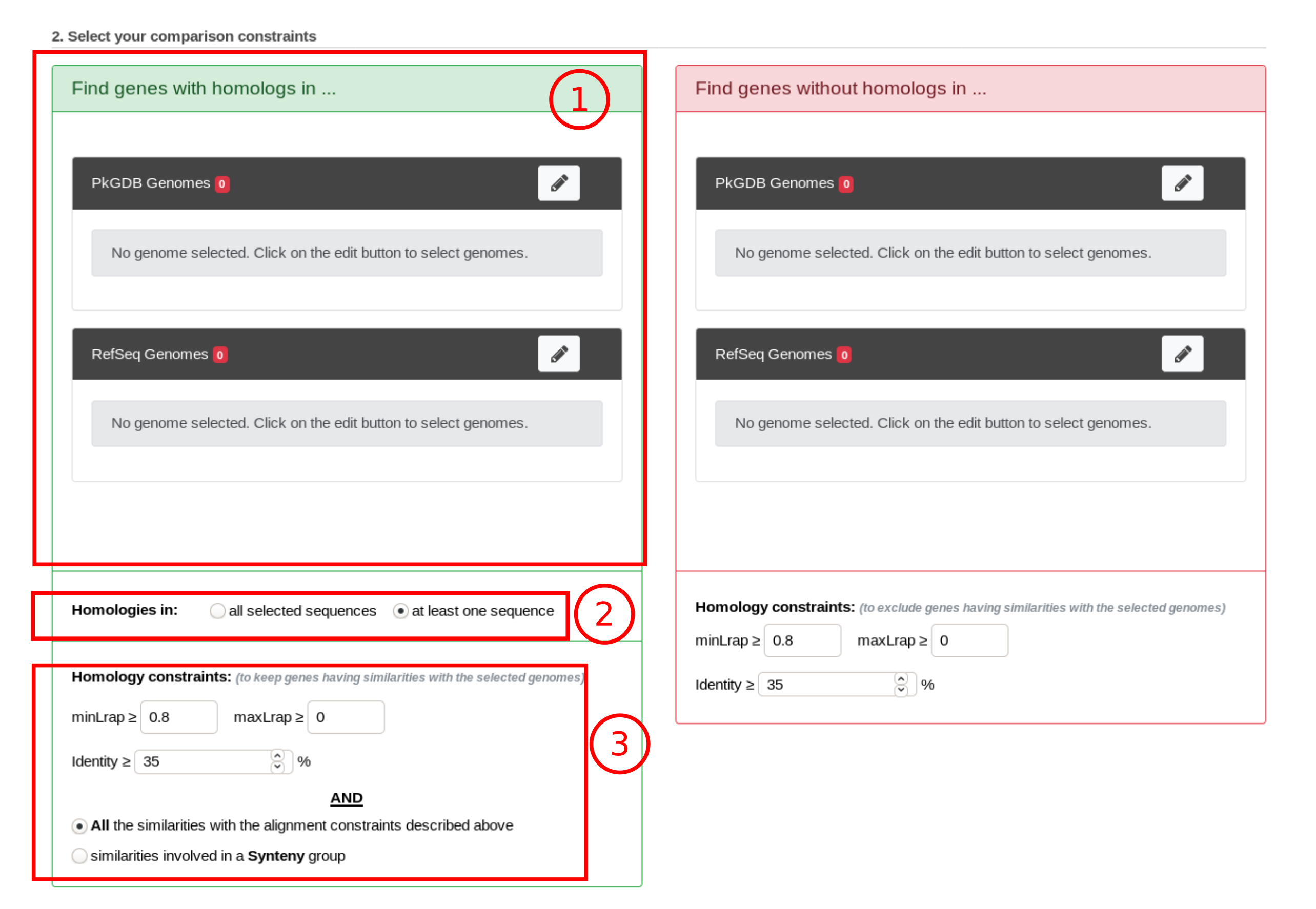
item C: Use this form to search for genes in your reference genome which have homologs in other organisms/replicons coming from PkGDB and/or RefSeq databases.
item D: Use this form to search for specific genes in your reference genome compared to a selection of organisms/replicons coming from PkGDB and/or RefSeq databases.
Forms C and D use the advanced selector (in Genome Selection mode). See here for help on how to use it.
Tip
You can mix the use of item C and item D to perform a very sensitive search. For example: get CDS of Acinetobacter baylyi ADP1 (reference genome, item A) which have homologs in Acinetobacter baumannii 6013113 and Acinetobacter baumannii AB0057 (item C), but NO homologs in Acinetobacter baumannii AYE (item D)
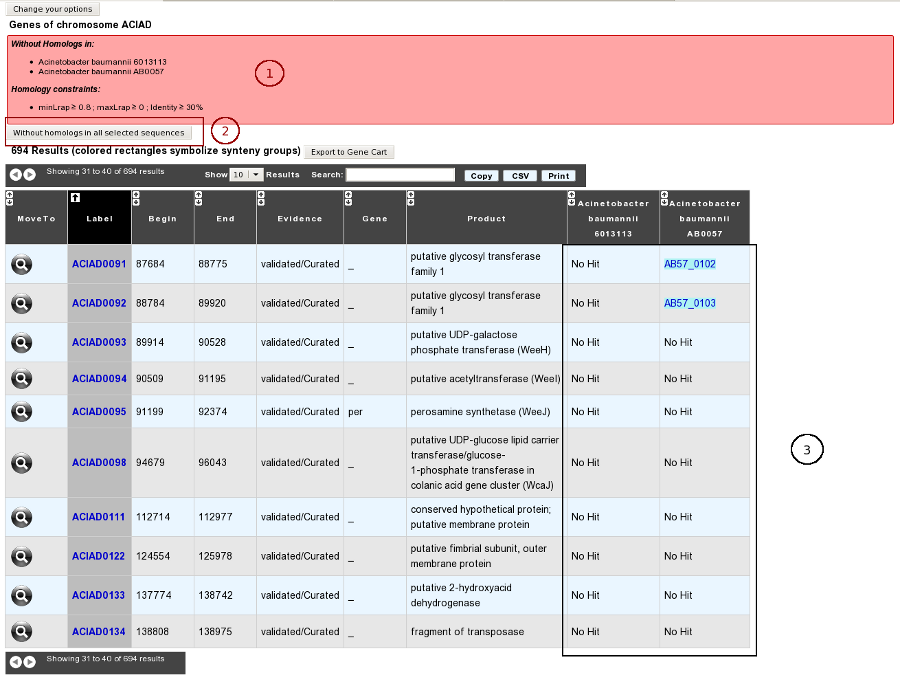
How to get genes with homologs in other organisms/replicons?
Results
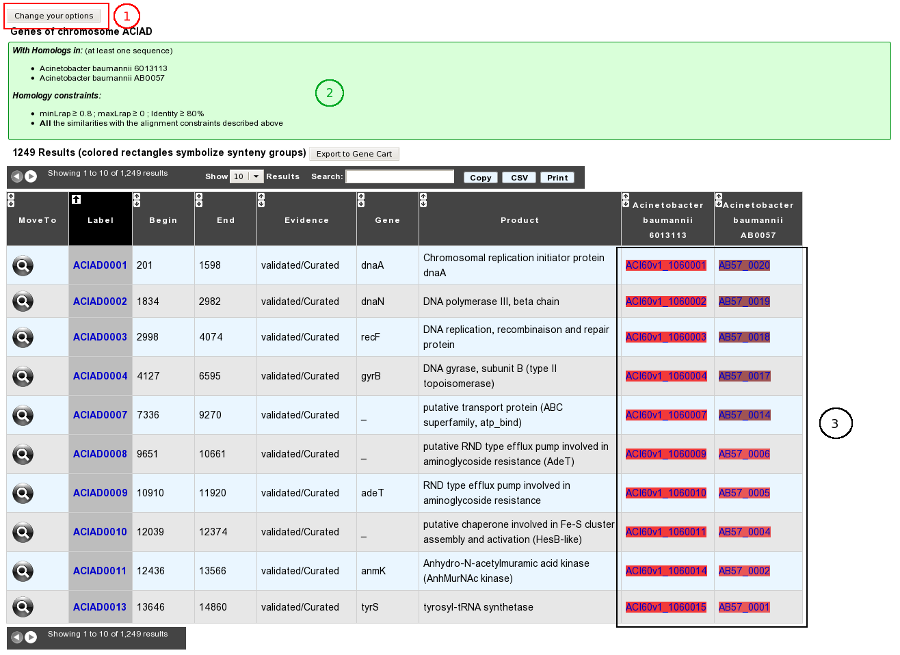
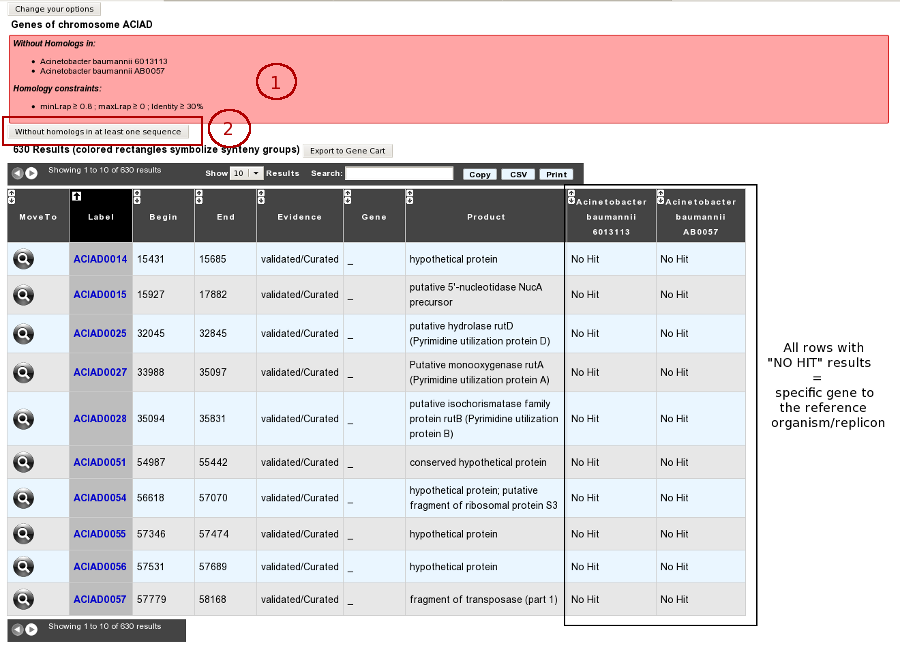
How to get specific genes of your reference genome compared to other organisms/replicons?
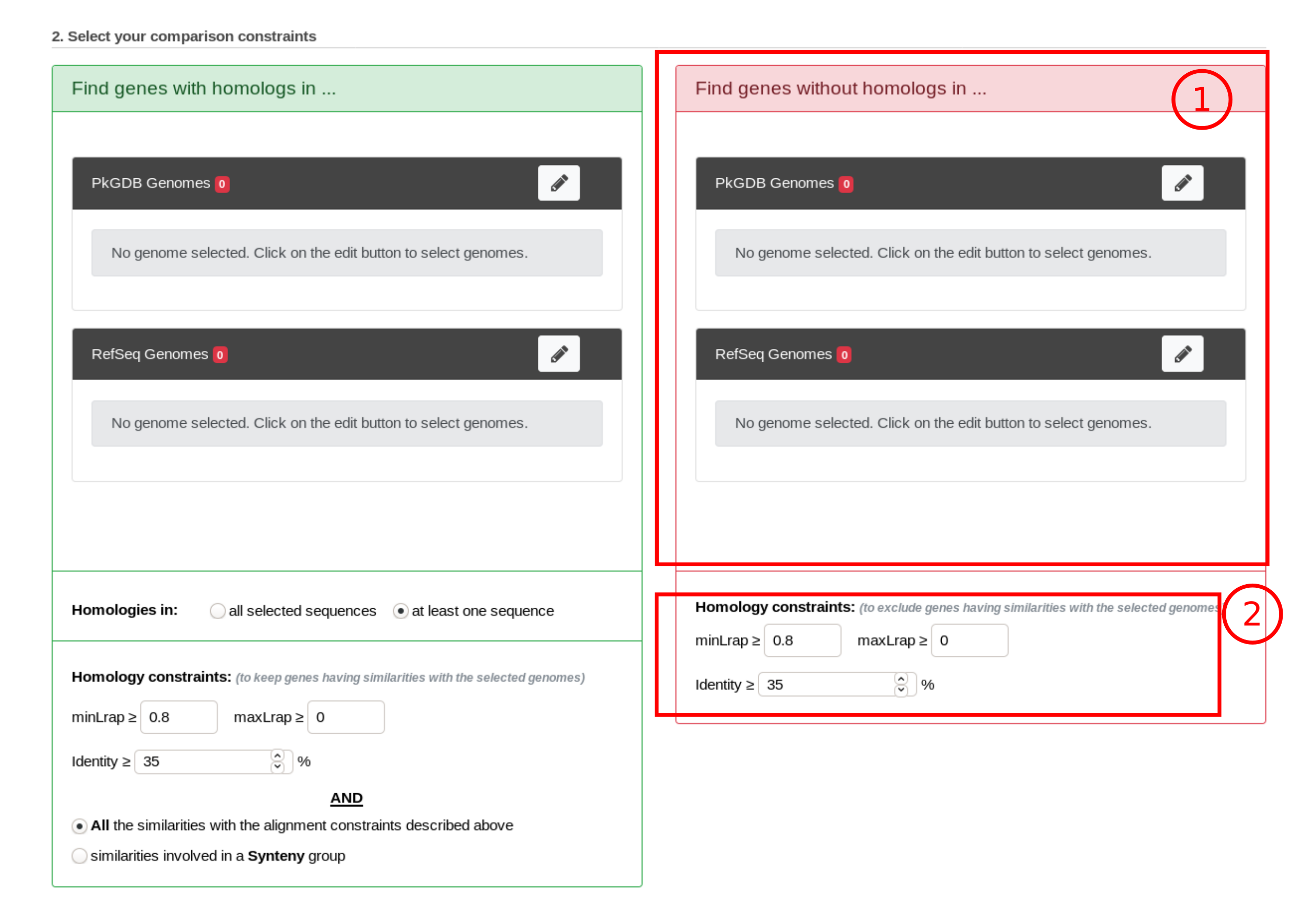
Results