Macromolecular Systems¶
What is MacSyFinder?¶
Macromolecular System Finder (MacSyFinder) provides a flexible framework to model the properties of molecular systems (cellular machinery or pathway) including their components, evolutionary associations with other systems and genetic architecture. Modelled features also include functional analogs, and the multiple uses of a same component by different systems. Models are used to search for molecular systems in complete genomes or in unstructured data like metagenomes. The components of the systems are searched by sequence similarity using Hidden Markov model (HMM) protein profiles. The assignment of hits to a given system is decided based on compliance with the content and organization of the system model.
Know more about MacSyFinder
Reference:
What type of Macromolecular systems can be detected?¶
MacSyFinder can detect :
- CRISPR-Cas systems: Clustered regularly interspaced short palindromic repeats (CRISPR) arrays and their associated Cas (CRISPR-associated) proteins form the CRISPR-Cas system. CRISPR-Cas are sophisticated adaptive immune systems that rely on small RNAs for sequence-specific targeting of foreign nucleic acids such as viruses and plasmids.
- a broad range of secretion systems: T1SS, T2SS,T3SS,T4SS, T5SS, T6SS, T9SS, Flg, T4P, Tad (Abby SS et al., Sci. Rep. 2016)
How to access to MacSyFinder predictions?¶
MacSyFinder predictions are available through the Comparative Genomics section, in the main navigation menu.
What is the ‘Macromolecular Systems’ table?¶
This table enumerates all macromolecular systems predicted for the selected organism and its replicons.
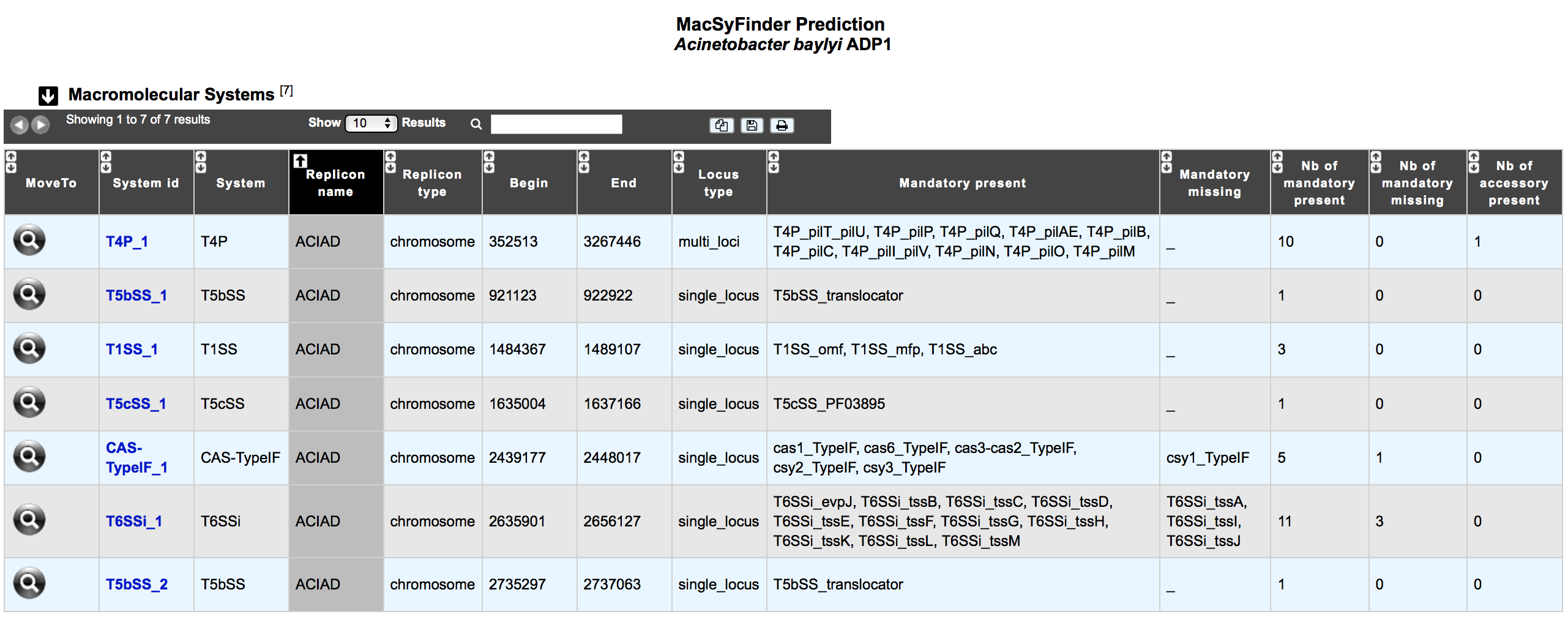
- System id: identifier of the system in the organism
- System: type of system detected by MacSyFinder
- Replicon name: identification of the replicon
- Replicon type: chromosome, plasmid or WGS
- Begin / End: Position of the system on the replicon
- Locus type: single or multi locus
- Mandatory present: list of mandatory genes of the system identified in the organism
- Mandatory missing: ist of mandatory genes of the system not detected in the organism
- Nb of mandatory present: number of mandatory genes of the system identified in the organism
- Nb of mandatory missing: number of mandatory genes of the system not detected in the organism
- Nb of accessory present: number of accessory genes of the system identified in the organism
How to explore a Macromolecular System?¶
The MacSyFinder System visualization window can be accessed by clicking on any cluster number in the System Id field. This window allows you to access to a detailled description of a selected Macromolecular System.